Group Wolinski
CELL BIOLOGY OF LIPID DROPLETS
Our research interests focus on the biogenesis and interactions of lipid droplets with other subcellular compartments. These universal organelles, from yeast to man, show multiple functions and play a critical role in lipid metabolism and energy homeostasis. Their dysfunction has been linked to different human diseases. We are especially interested in the spatio-temporal mechanisms of lipid droplet formation and maintenance at the molecular and cellular levels. Our studies involve molecular, cell biological, and analytical techniques. For cytological studies, we apply state-of-the-art bioimaging systems such as confocal laser scanning microscopy, selective plane illumination microscopy, CARS microscopy, or superresolution technologies (STORM). In addition, we develop and integrate advanced image-informatics methods for quantitatively assessing lipid droplet dynamics. We use genetically modified yeast model systems and mammalian cell systems to better understand the mechanistic basis of lipid-droplet-associated disorders in humans.
Role of seipin in local phosphatidic acid homeostasis
Berardinelli–Seip congenital lipodystrophy type 2 gene (BSCL2) encodes the protein BSCL2 (“seipin”). Loss-of-function mutations in BSCL2 are associated with congenital generalized lipodystrophy type 2 (CGL-type 2), the most severe genetic lipodystrophy disease in humans leading to an almost complete absence of adipose tissue (Ref). CGL-type 2 is unhealable. Therapeutic options are limited to the treatment of occurring metabolic syndromes. Seipin function is studied for decades in different cell systems including the yeast Saccharomyces cerevisiae. However, the specific molecular function(s) of seipin and its role in the pathogenesis of CGL-type 2 is still elusive and controversially discussed.
At the cellular level, seipin malfunction affects both neutral lipid and glycerophospholipid metabolism exemplified in yeast, mice, other model systems and human cells by multiple abnormal cellular phenotypes such as aberrant formation and assembly of neutral lipids stores, altered phospholipid membrane synthesis and distribution, or aberrant ER and mitochondrial calcium homeostasis.
We hypothesize that seipin mainly serves as a lipid-binding protein in particular for phosphatidic acid and other anionic lipids, besides its role in LD assembly. Lack of seipin leads to locally elavated PA levels at LD-ER contacts sites and at an LD-forming subdomain of the nER, causing different abnormal cellular phenotypes in yeast (Wolinski* et al. 2015. BBA Molecular and Cell Biology of Lipids).
Using cell and molecular biology methods in combination with advanced optical imaging technologies available at the IMB-Graz (Optical Imaging Resource) we aim at differentiating direct from indirect spatio-temporal effects caused by seipin malfunction and different experimantal setups to uncover the trigger(s) causing the disease in humans.
K. Skrobar, S. Mandl, H. Wolinski
Quantitative optical imaging to probe lipid droplet biology
Quantification of multi-dimensional image data obtained from microscopic techniques such as confocal laser scanning microscopy has become a fundamental approach to probe cellular events at the single-cell level. However, image-based quantification is still a challenge and error-prone (Pribasnig et al. & Wolinski 2018*. BBA Molecular and Cell Biology of Lipids). We are currently evaluating different strategies for more reliable quantiative analyses particularly of lipid droplets and associated structures/organelles of the yeast, Saccharomyces cerevisiae. In this context, we use a variety of open-source software tools available on different platforms such as Matlab, Phyton, or Fiji/ImageJ including machine learning methods for cell detection.
K. Skrobar, H. Wolinski
Probing LD dynamics and heterogeneity using photoconvertible fusion proteins and live cell STORM
Yeast lipid droplets are partially inherited to daughter cells during cellular growth leading to lipid droplet subpopulations (inherited - newly synthesized; Wolinski et al. 2009. JCS; Wolinski* et al. 2015. BBA Molecular and Cell Biology of Lipids). Currently, it is not possible to probe selected subpopulations of lipid droplets. We are evaluating photoconvertible fusion proteins and developing methods for selective conversion of a fraction of lipid droplets to address lipid droplet heterogeneity. Furthermore, we establish superresolution live cell STORM for detailed localization of proteins involved in phospholipid and neutral lipid metabolism including seipin. In addition, we apply FRAP to study the dynamic behavior of a subset of proteins involved in lipid droplet biogenesis and degradation to analyse protein crowding during these processes.
H. Wolinski
Quantification of neuronal structures in whole-mount cleared adipose tissue
Recent studies highlighted the importance of neuronal innervation of (white) adipose tissue for the metabolic regulation of adipocytes. Optical clearing and high-resolution imaging technologies are the methods of choice to study the organization and structure of adipose tissue under different experimental conditions. However, quantification of the complex neuronal network in large tissue samples is still challenging. In this collaborative work (Working group Schweiger), we focus on developing flexible image-informatics pipelines for the accurate quantification of neurons in large 3D data sets. Our work includes the development of benchmark systems for testing different segmentation and quantification approaches. In this respect, pitfalls and limitations of existing procedures for the quantification of neurons are evaluated. Importantly, our studies are based on the analysis of entire adipose tissue.
Rauchenwald, Schweiger, Wolinski.
Previous work
Lipid droplet interactions revisited:
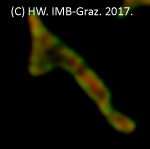
Pribasnig, M., Kien B., Pusch L., Haemmerle G., Zimmermann, R. & Wolinski H. (2018). Extended-resolution confocal imaging of the interaction between lipid droplets and mitochondria. BBA Molecular and Cell Biology of Lipids, 1863(10):1285-1296
Movie: 'Fusion and fission' of mitochondrial structures in COS-7 cells monitored using extended-resolution 4D live cell imaging.