Research
Design and functionalization of new nanodiscs
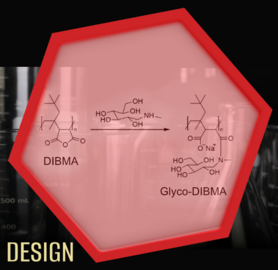
A few years ago, amphiphilic copolymers were introduced as an alternative to traditional detergents. These polymers wrap around phospholipid bilayers like a belt, creating nanodiscs that enclose membrane proteins. We established a diisobutylene/maleic acid (DIBMA) copolymer as a new protein-extracting agent that offers key advantages over other polymers: The ordering of lipid chains embedded in nanodiscs is only slightly affected by DIBMA, and spectroscopic studies of membrane proteins are possible also in the important ultraviolet spectral range. In addition, DIBMA tolerates elevated concentrations of divalent cations without precipitating, which is important for activity studies of many membrane proteins. We further developed a polymer derivative called Glyco-DIBMA that retains the same favorable properties but allows even more efficient solubilization of lipids and extraction of membrane proteins. Recently, we have shown that even some small molecules, such as compounds with a diglucose head group, are able to extract membrane proteins and their surrounding lipids directly into nanodiscs, making them accessible to detailed studies.
We are designing new polymers and small-molecule compounds to further improve nanodiscs for membrane proteins, making them even more compatible with biophysical and structural biology applications.
Biophysics of membrane proteins in nanodiscs
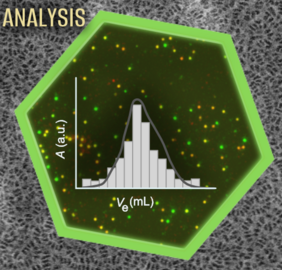
As a second focus, we employ a wide range of biophysical methods to study nanodiscs and the membrane proteins and lipids embedded within them. Our biophysical methods includes numerous spectroscopic, calorimetric, and chromatographic techniques as well as light scattering and microfluidic diffusional sizing. In addition, high-resolution imaging techniques such as electron microscopy and single-molecule fluorescence measurements are used, as well as high-throughput technologies such as mass spectrometry for proteomics and lipidomics. These complementary methods provide detailed information on the structure and size of nanodiscs as well as the conformation and dynamics of membrane proteins and lipids.
We are continuously refining these methods to make them more compatible with nanodiscs and, conversely, use the gained insights for the rational optimization of nanodiscs.
Nanodiscs as platform for biomolecular applications

A particularly important aspect of our research is the use of new nanodiscs as versatile tools to answer biomolecular questions. Nanodiscs allow both the isolation of individual proteins and protein complexes as well as the extraction of entire protein libraries from different organisms and cells. This makes it possible to perform functional and interaction studies under laboratory conditions that can be well controlled, but still mimic the natural environment of membrane proteins. For example, using the above-mentioned methods, protein/protein interactions can be studied within a membrane environment without the need to purify or reconstitute membrane proteins. Our nanodiscs are also employed for high-resolution structural analyses of membrane proteins by electron microscopy, and can be used for ligand binding studies, thereby facilitating the identification of new drugs.
We are developing this nanodisc technology platform to elucidate the structures, dynamics, and interactions of membrane proteins with high resolution yet in an environment similar to the natural membrane.
Some of our polymers are available from GLYCON.


